 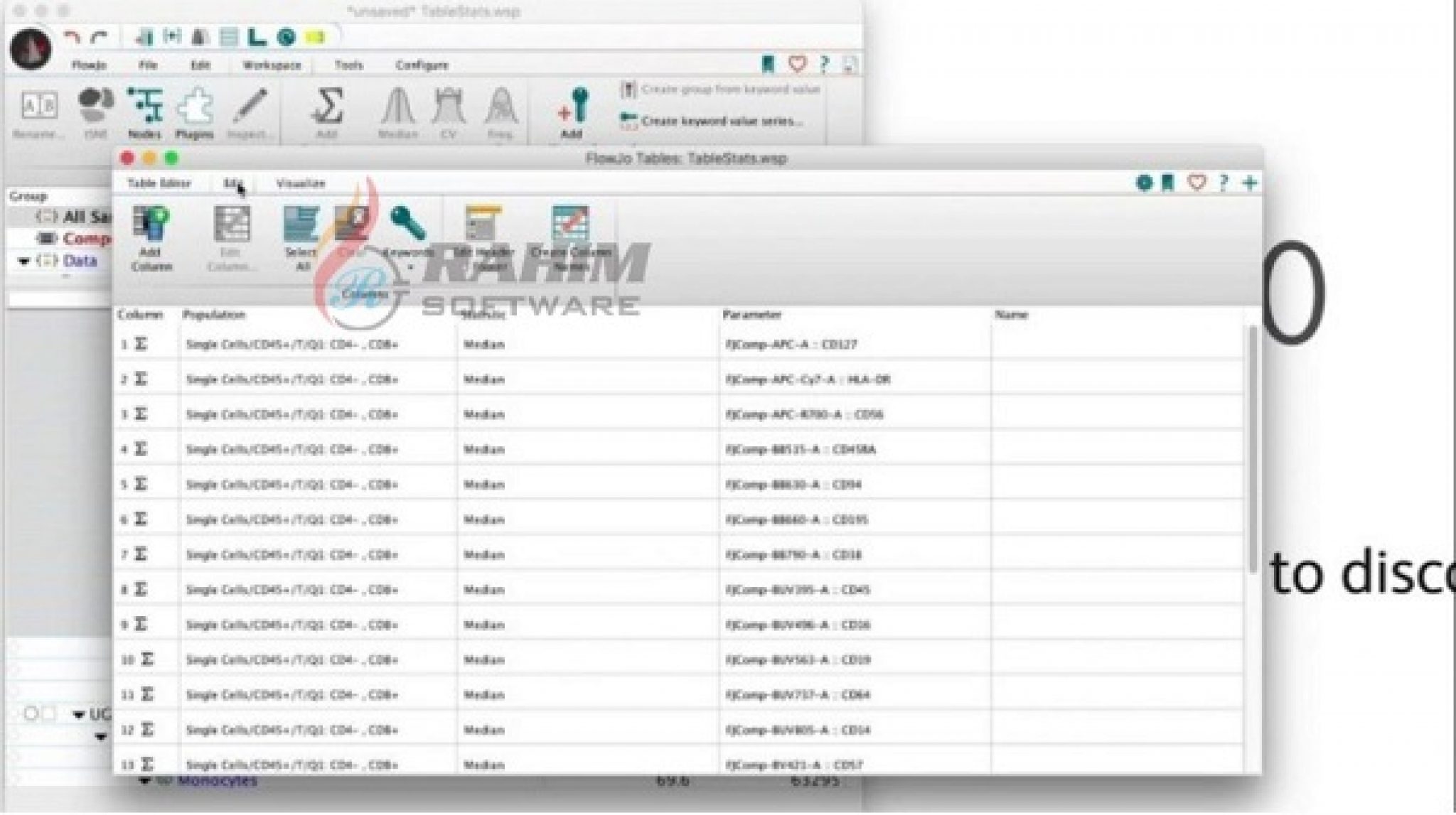 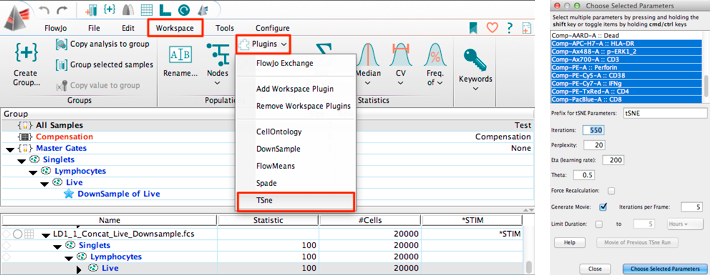
To actually use this data elsewhere (such as with R), export it into a csv format which can be easily imported by other programs. This will make a new window, holding the table you wanted. With the settings fixed, you can hit the “Create Table” button at the top of the main workspace. To do this, double click on that row, and select the statistic you want to include.Īnd you should now have something that looks like this: For example, I find it much more informative to have the singlets show total count, rather than Freq of parent. Some of those statistics aren’t what we’re looking for. Once in this window, select the populations / statistics you want to include from the main workspace, and drag it into the table editor, so you have something that looks like this. Lastly, the easiest way to output this data is to hit the Table Editor button near the top of the screen to open up a new window. Of course, a scatterplot needs a second axis, so I just used mCherry fluorescence (or the lack of it, since these were just normal 293T cells), captured by the YL2-A detector.Īnd of course copy that to the group as well (you should know how to do this by now). While this can be done in histogram format, I generally also do this with a scatterplot, since it allows me to see even small numbers of events (which would be smashed against the bottom of the plot if it were a smoothed histogram). In this case, it’s GFP positivity, captured by the BL1-A detector. Next is actually setting up the analysis for the response variable we were looking to measure. Then only gate the cells directly on the diagonal, thus excluding those that have more FSC-A relative to FSC-H. To do this, make a scatterplot where FSC-A is on the X-axis, and FSC-H is on the Y-axis. Here is the best description of doublet gating I’ve seen to date. So the live cell population will already be enriched for singlets, but having a second “doublet gating” step will make it that much more pure. Once you do that, the population should now be in bold, with the same text color as the group name. Thus, right click on the “Live” population in the workspace and hit “Copy to Group”. Once you have made that gate, you’ll want to keep it constant between samples. Since we’re mostly using HEK cells, that means that main thing we will be doing in this screen is gating for the population of cells while excluding debris (small FSC-A but high SSC-A). Set it so forward scatter (FSC-A) is on the X-axis, and side scatter (SSC-A) is on the Y-axis. By default, a scatterplot should show up. the first sample), and double clicking on it. Start by choosing a representative sample (eg. Now to actually start analyzing the flow data. 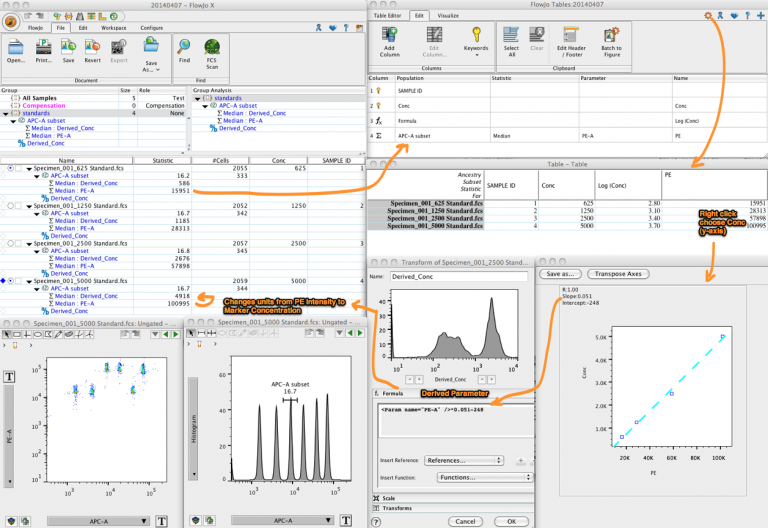
Now that the group is made, I select it, and then drag the new sample files into it, like below: Thus, I hit the “Create Group” button and type in the name of the group I’ll be analyzing. Before you start dragging in samples, I find it useful to make a group for the specific set of samples you may want to analyze. With that out of the way, now you can perform your analysis. Thus, click on the “Preferences” button:Īnd finally once in that final window, change the “Decimal Precision” value to something like 8. This can be prohibitively uninformative if you have very low percentages that you’re trying to accurate quantitate. Once logged in, you’ll be starting with a blank analysis workspace, as below.īefore I forget, an annoying default setting of FlowJo is that it only lists two decimal points in most of its values. I’m obviously not going to share my login and password here ask someone in the lab or consult the lab google docs. Obviously, if you don’t have FlowJo yet, then download it from the website. Thus, here’s a short set of instructions for using it to do a basic process, such as determining what percentage of live cells are also GFP positive. While I’ve been using FlowJo for a long time, I realize it isn’t super intuitive and new people to the lab may first struggle in using it. Inevitably, what ends up being the most practical tool for analysis of low cytometry data is FlowJo. We do a lot of flow cytometry in the lab. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
